Customizing the game
Often people ask how to install a moviepack, how you make your cycle color different from your tail's, etc. Read on if you want to know how.
Moviepacks
Installing a Moviepack
Moviepacks are packages that completely change the look of the game. the floor, the walls, the cycles, everything. Nowadays, you can download many from different places around the net; for pointers, consult the Moviepacks list. They usually come in a ZIP file containing a moviepack folder.
This folder goes into your ArmagetronAdvanced folder (for Windows users usually in C:\program files\armagetronadvanced\). If this moviepack folder doesn't exist yet, and you don't happen to find it in the archive you've downloaded, you can create one by yourself.
Default Armagetron Advanced installation folder for system wide installation
Windows: C:\Program Files\Armagetron Advanced\
Unix/Linux: /usr/local/share/games/armagetronad/ or /usr/share/games/armagetronad/
Mac OS X: The game is self-contained and may be located anywhere, but most people usually have it in /Applications
The moviepack folder (from .zip file) can be placed inside the "Armagetron Advanced.app" package bundle: The moviepack folder (extracted from .zip file) should be placed in Contents/Resources (click on "Armagetron Advanced.app" in Finder then Click Action Icon (looks like Gear) and choose "Show Package Contents" to open package bundle)
Default Armagetron Advanced User Data Folder for user specific installation (recommended)
- Note that you have to enable viewing hidden files to see some directories.
- For all systems, unpacking the moviepack so that it sits as a 'moviepack' folder inside the user data folder works.
Windows 2000/XP: C:\Documents and Settings\[Username]\Application Data\Armagetron
Windows Vista: C:\Users\[Username]\AppData\Roaming\Armagetron
Unix/Linux: ~/.armagetronad
Mac OS X: ~/Library/Application Support/Armagetron Advanced
TODO: put folder locations in a single wiki entry and validate
The armagetron installation and configuration paths should be somewhere else on the wiki, too
Problems
Windows
Sometimes you get errors after installing a moviepack, such as
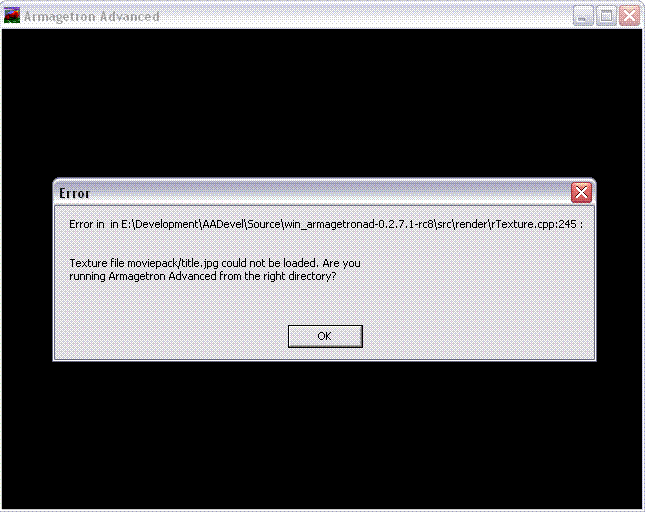
this means that the moviepack you installed is not complete, i.e. Armagetron Advanced looks for files that are not present in the moviepack. The easiest way around this is to first install the original moviepack (get the ZIP) and install the moviepack you actually want on top of it.
Unix/Linux
In Unix based Operating Systems, where file case is important, sometimes a texture doesn't display even though it appears to be installed correctly. to correct this, manually change the case of the file to suit. In particular, upper case extensions can cause problems. ie, change filename.JPG to filename.jpg and it will probably be OK. If settings.cfg is partially or completely capitalized, no textures will load even if they are properly capitalized. You should inform the author of the moviepack of the problem so he can fix it.
Installing a new model
In your Armagetron Advanced moviepack directory (see above) locate bike.png and cycle.ase and replace with the new ones. It is important to note that there must be an existing moviepack installed to single update to a new model.
Customizing default textures
Installing a different default texture
ArmagetronAd allows to change textures of the floor, the walls, the cycles and the sky. All textures are contained in the "textures" folder. Textures are images displayed during the game. For exmaple the image on the outer wall can be changed by exchanging the rim_wall.png file. Instead of replacing any texture files, you should rename the original file and then copy the new one into the folder. Now you have a backup available if you want to restore the original.
Where to find those textures ?
Windows: C:\Program Files\Armagetron Advanced\
Unix/Linux: /usr/local/share/games/armagetronad/
Mac OS X: ~/Library/Application Support/Armagetron Advanced/
Simply drag a moviepack folder to this location. Folder must be named "moviepack" only (no extensions or numbers).
'TODO: put folder locations in a single wiki entry and validate'
Which file is what ?
rim_wall.png = texture for the outer walls
sky.png = texture for the lower sky (the upper sky uses the floor*.png)
floor.png = single textured floor (and upper sky)
floor_a.png, floor_b.png = dual textured floor
bike.png = cycle's texture
Creating a new texture
Creating a texture is as simple as making an image and saving it as described above. For the best results, follow these guidelines:
Power-of-two sizes
The size of OpenGL's standard 2D textures must be a power of two. If the size of an image isn't, the game (or more accurately, GLU) will scale it to the nearest power of two before sending it to OpenGL, resulting in lower quality. Powers of two include:
- 2^0 = 1
- 2^1 = 2
- 2^2 = 4
- 2^3 = 8
- 2^4 = 16
- 2^5 = 32
- 2^6 = 64
- 2^7 = 128
- 2^8 = 256
- 2^9 = 512
- 2^10 = 1024
- 2^11 = 2048
Note that a texture need not be square; 512x64 textures are fine.
Use transparency wisely
There are several things to keep in mind when using transparent textures:
- Because of to the way OpenGL prevents background objects from drawing over foreground objects, it might not be possible to see everything through a transparent texture.
- Colors will bleed from neighboring texels (texture pixels) if both color and transparency are different. An exception to this is the cycle texture, which is preprocessed by the game and has no transparency in the end.
Cycle Colors
The basics of cycle color
First you have to understand how computers understand colors. In armagetron, colors are saved with three values: the red, green and blue (R, G and B) parts. Each of these three so-called RGB values can hold a number from 0 to 15 - 0 means none and 15 means as much as possible. A few examples:
r 15, g 0, b 0 : strong red r 0, g 15, b 0 : strong green r 0, g 0, b 15 : strong blue r 8, g 0, b 0 : darker red
you can also combine the colours (follows the same rules as when you combine red,green and blue light):
r 15 g 0 b 15 : strong pink r 0 g 15 b 10 : strong cyan r 15 g 15 b 0 : yellow r 15 g 8 b 0 : orange (less green => more red than yellow)
you get the gist. So how do you set these colours ? you can set them via the player setup menu, but if you want to have different cycle and trail colours, you have to use the Console. The commands are
color_r_1 X # set red component of the 1st player's cycle to X color_g_1 Y # set green component of the 1st player's cycle to Y color_b_1 Z # set blue component of the 1st player's cycle to Z
The best way to learn how to write the colour you want is to experiment a bit.
Different Cycle and trail colors
Remember me saying that the colors were saved as numbers between 0 and 15 ? Well, that was not entirely true. If you set a color to a value over 15, the trail part is set to 15 and the cycle is (usually) set to the actual value minus 16. For example:
r 16 g 16 b 0 : yellow tail (r 15 g 15) and black bike (16 - 16 = 0)
higher numbers can give you multi-colored trails. Again, experiment.
For an exact explanation, see elaborate color explanation.
Developer's note: this behavior of the game is technically a bug and is officially unsupported. It is tolerated, though, because it's fun :) But some servers, especially team play servers where your color is no mere decoration, will clamp your color settings to 15 and this trick does not work.
Also check out this useful link for helping you in picking colors:
http://armagetron.co.uk/colours/colourpicker.php
Chat Colors
If you want to use different colors when chatting you have to use a different syntax for colors. For example if you want to say a text in red you would enter (at the chat prompt):
0xff0000 this appears in red
note that this is all zeroes (not o's), so zero-xff-zero-zero-zero-zero. This is important since it won't work otherwise.
The first two digits after the "0x" indicate the red part of the color, the two digits after that the green one and the two last ones the blue part. They signify numbers from 00 to ff (0 to 255) written in hex (base 16); people who ever used color codes in HTML or CSS will be familiar with this. Only lowercase letters are accepted in the code.
A few different two- letter combos are as follows:
- ff: works like 15 in the bike color
- ee: like 14
- dd: 13
- cc: 12
- bb: 11
- aa: 10
- 99: 9
- 88: 8
- ...
- 00: 0
Note that you can use different hex numbers like in 0xab1234 which lets you chose your color more exactly, but in most cases you won't need this. If you are so paranoid that you think you need exactly your shade of pink feel free to read the article Bases on this.
Other combos to get you started are:
0x00ff00 this is green 0x0000ff this is blue 0xff8800 this is orange 0xffffff this is white 0xffff7f this appears in the normal chat yellow (no change)
Developer's note: these color codes were not intended for use by the players; the choice of 0x as intro string was a bit unfortunate. Some servers will filter away your codes, so do your testing in a local game. Since it was not intended for use, no thought was given on usability, and for example upper case codes are not accepted. So use 0xff0000 for red, not 0xFF0000. And don't be angry about future unannounced changes to the format.
Autoexec.cfg
As you can tell from above, Armagetron is insanely customizable. But what if you want to make semi-permanent changes to settings you might type into the Console? The answer is the autoexec.cfg file. Here is how it use it:
1. Locate the configuration directories. Launch Armagetron and select "About" from the main menu. This will list directories where Arma stores/reads data. The configuration directory you want access to is usually in your $HOME folder (this is dependent on your operating system). Open and bookmark/shortcut the configuration directory, you'll probably want to come back here later. On Arma v0.4 and greater the path is “Help” -> “Open Directories” -> “Open User Configuration”.
2. Create or edit autoexec.cfg. If this file does not exist, create it. There is nothing special about CFG files, they can be edited with any simple text editor like Notepad on Windows. Armagetron looks for and loads this file when it starts. Any settings you put in there will override default values.
3. Add your custom settings. Some of the most common setting are for Custom Camera (rise, pitch, etc.). You can see a list of these and many, many more on the Console Commands page. Put each setting on it's own line followed by a space, then the value of your choice. Save the file when finished.
4. More settings. autoexec.cfg will also override settings in your user.cfg. You can copy instant chats, your player name, even keybindings found in user.cfg to autoexec.cfg and modify them without breaking your user.cfg file.
5. Advanced usage. You can have any number of config files in the directory alongside of autoexec.cfg, with any name you choose. This is helpful if you want a modular setup. For instance, you can store camera settings in a file named "my_camera.cfg" and dual cycle colors in "my_cycle.cfg", then use autoexec.cfg to load them with the INCLUDE command. Example:
INCLUDE my_camera.cfg INCLUDE my_cycle.cfg
6. Even more advanced usage. Configurations can be loaded at any time, not just at start. Maybe you have texture settings and instant chats that are specific to a server such as Fortress. Let's say it's called fortress.cfg. You can enter chat and type: /console include fortress.cfg to load those settings. To make things easier and faster, set an instant chat key to /console include \\ and type the filename. (Remember, loading custom settings in this way will override others, so you probably want to save your custom default values in autoexec.cfg or make a reset.cfg file to reverse the changes.)